|
3/16/2023 0 Comments Snapgene viewer pdest 17 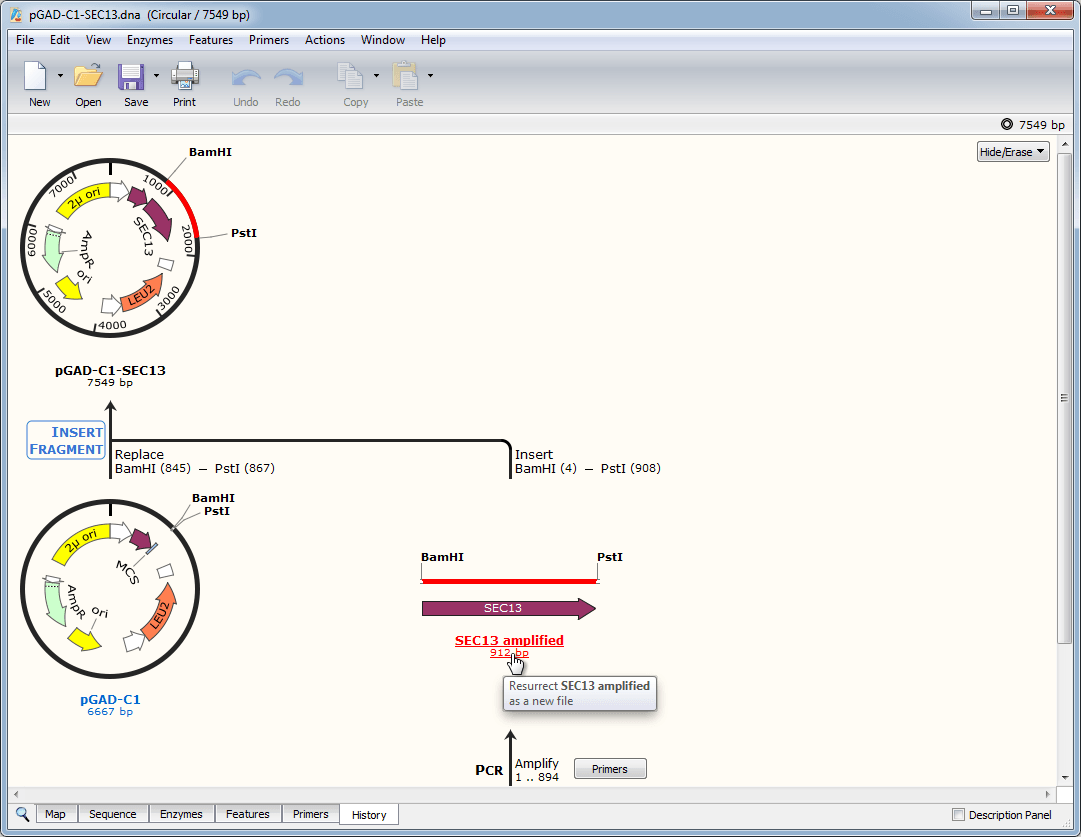
In this report, we probed this specific peptide binding function of AGR2 to define a possible biological function for this activity. EpCAM forms a model client protein for AGR2 to study how an ER-resident chaperone can dock specifically to a peptide motif and regulate the trafficking a protein destined for the secretory pathway. These data define a dominant site on AGR2 that mediates its specific peptide-binding function. Hydrogen-deuterium exchange mass spectrometry was used to identify a stable binding site for AGR2 on EpCAM, adjacent to the TLIYY motif and surrounding EpCAM's detergent binding site. Introducing a single alanine mutation in EpCAM at Tyr251 attenuated its binding to AGR2 in vitro and in cells. Proximity ligation assays demonstrated that endogenous AGR2 and EpCAM protein associate in cells. AGR2 and EpCAM proteins formed a dose-dependent protein-protein interaction in vitro. Screening the human proteome for proteins harboring this motif revealed an enrichment in transmembrane proteins and we focused on validating EpCAM as a potential AGR2-interacting protein. A peptide binding site consensus of Tx was developed for AGR2 by measuring its activity against a mutant peptide library. Hydrogen-deuterium exchange mass spectrometry localized the dominant region in AGR2 that interacts with the TTIYY peptide to within a structural loop from amino acids 131–135 (VDPSL). A synthetic TTIYY-containing peptide column was used to affinity-purify AGR2 from crude lysates highlighting peptide selectivity in complex mixtures. AGR2 protein has a relatively unique property for a chaperone in that it can bind sequence-specifically to a specific peptide motif (TTIYY). AGR2 is an oncogenic endoplasmic reticulum (ER)-resident protein disulfide isomerase. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
